3 steps of translation that bring proteins for you
by egpat 17-05-2017
We are living every day and our cells continuously growing from our childhood. Your muscles develop, cells are dividing and many biochemical reactions in your body make you intact and highly functional. How it all can happen? Simply, proteins do the major job for us sometimes being structural and sometimes being functional such as enzymes. Its fine, are they need to be supplied in diet?
Of course, they are required in diet but not for supplying proteins directly into the body instead they simply act as a source of amino acids. From these amino acids, our body can synthesize proteins. This protein synthesis is guided by a process called translation that can prepare proteins form amino acids.
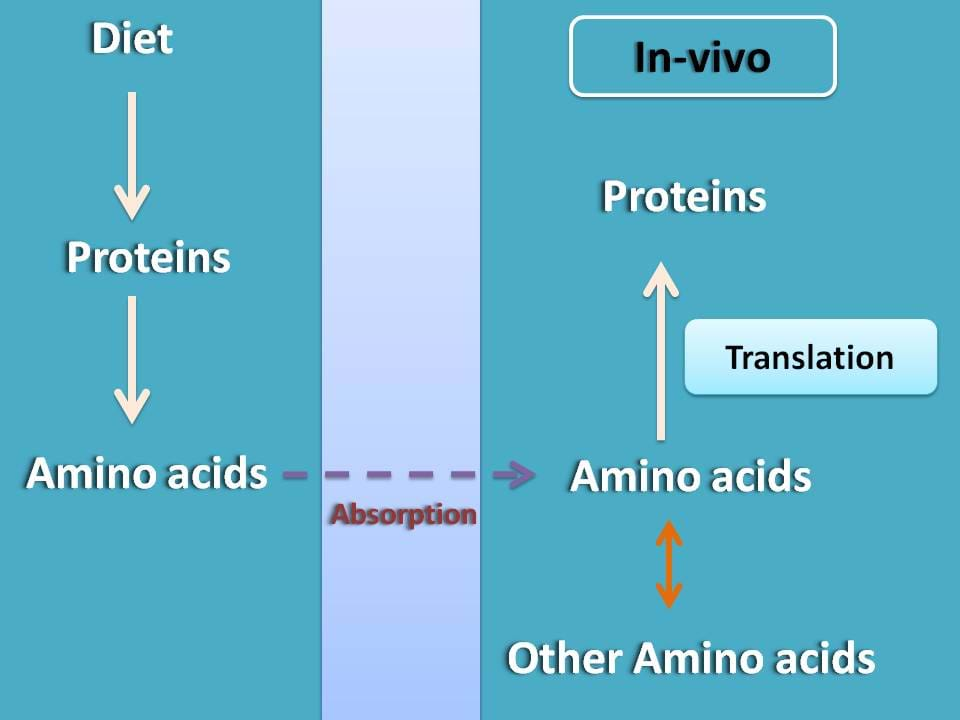
Few of the amino acids are synthesized within the body from amino acids supplied in diet by transamination. In a simple terms, translation is a process of protein synthesis from all these amino acids based on the code present in mRNA.
What you need for protein synthesis?
Mainly it requires three components each playing their unique role in protein synthesis.
mRNA
This is a piece of transcript that is derived from DNA and contains all the genetic information required for protein synthesis. It tells which amino acid should be there and how they are to be arranged by specific codon coding for each amino acid.
tRNA
Okay, Its fine that mRNA gives code for amino acid sequence, but who brings each amino acid and attach in such a sequence? That is none other than, tRNA, named a transfer RNA.
rRNA
rRNA is the ribosomal RNA that acts as work place for protein synthesis and responsible for holding mRNA and tRNA. In addition to these three components, translation also requires few of the proteins acting as eukaryotic activating factors which we will see in our further discussion.
With these basic things, now let’s see the various steps in translation. These overall process can be broadly categorized into three steps.
- Initiation
- Elongation
- Termination
Initiation
It is well known fact that ribosome will have two parts one being bigger and another one as smaller. These subunits in eukaryotes generally exist as 60S and 40S while in prokaryotes they exist as 50S and 30S.
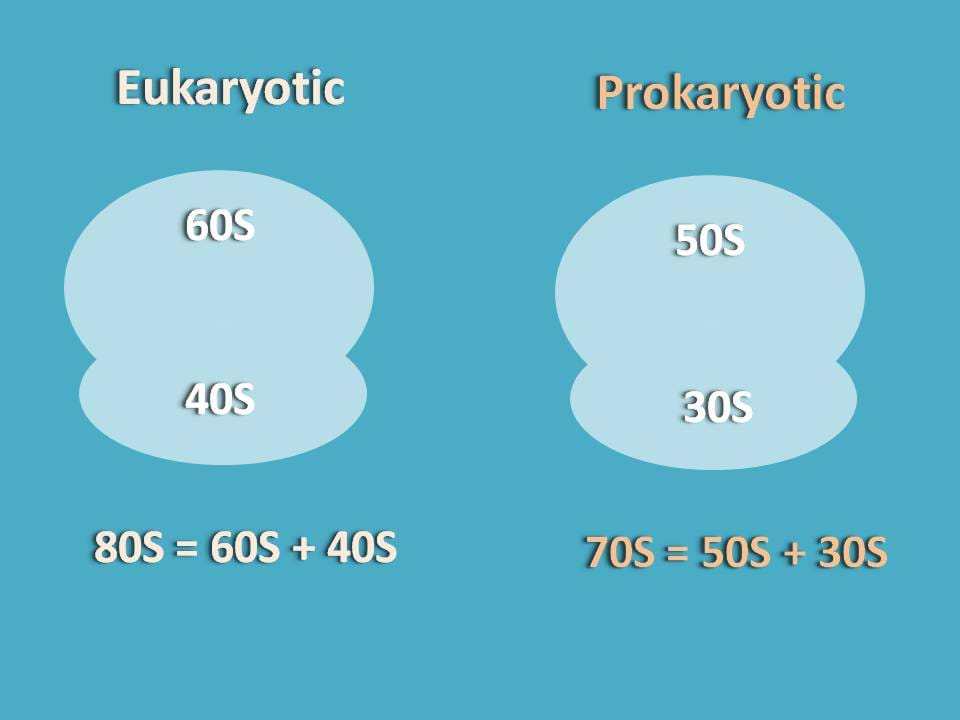

We will see here the translation of proteins in eukaryotes in detail.
So, our first question is that, to which part mRNA and tRNA bind?
Since mRNA is carrying genetic code, it binds to smaller part while tRNA bringing each amino acid binds to bigger part.
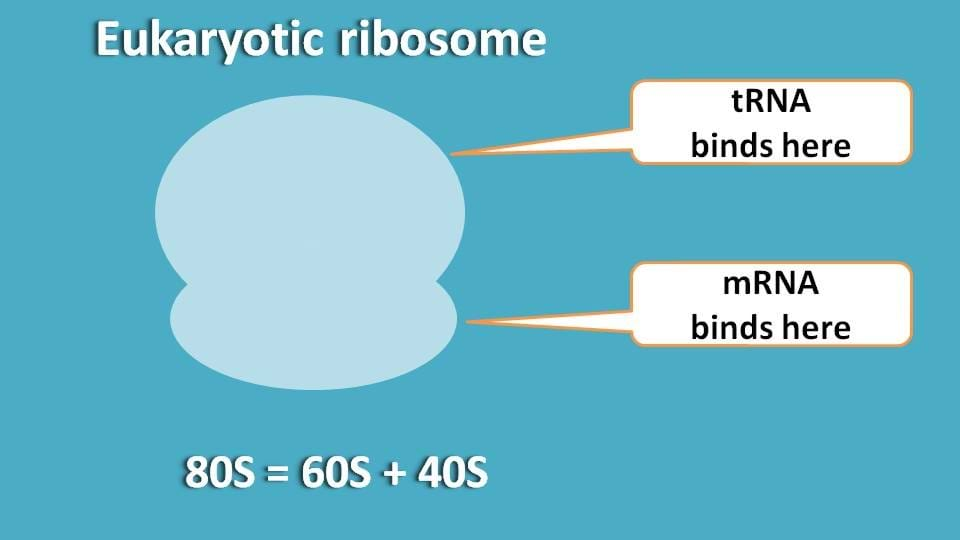
So, can you guess now which subunit is to be more modified to facilitate translation?
Undoubtedly, it is 40S subunit as this is the site where mRNA binds and brings all the genetic information required for translation.
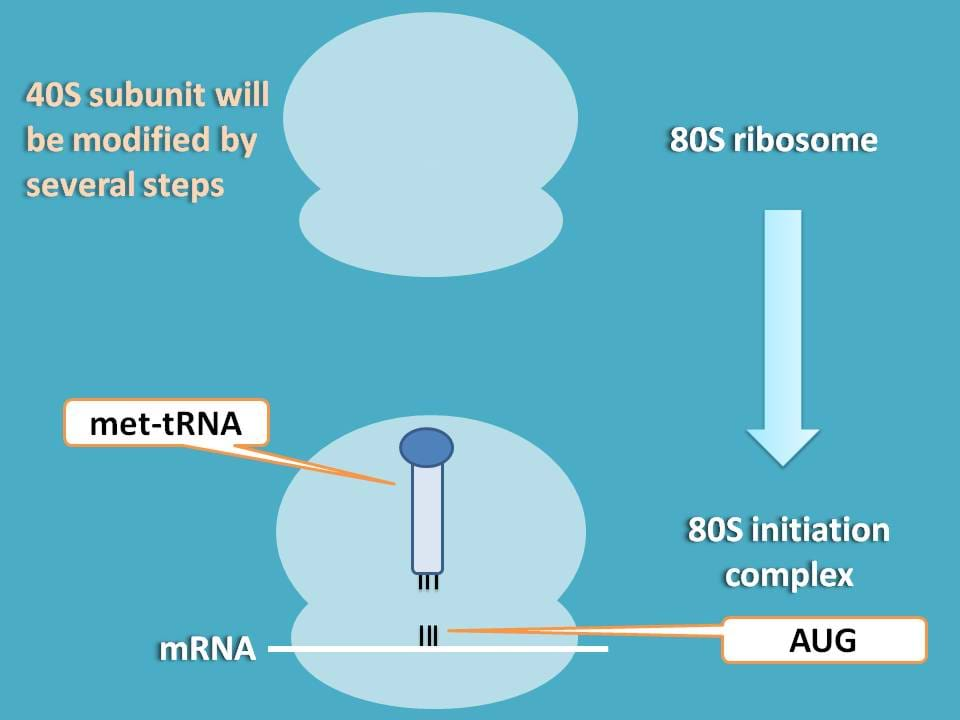
So, initiation of translation involves various steps all interlinked with one to another.
- Formation of 60S and 40S subunits
- 43S preinitiation complex
- 43S initiation complex
- 48S initiation complex
- 80S initiation complex
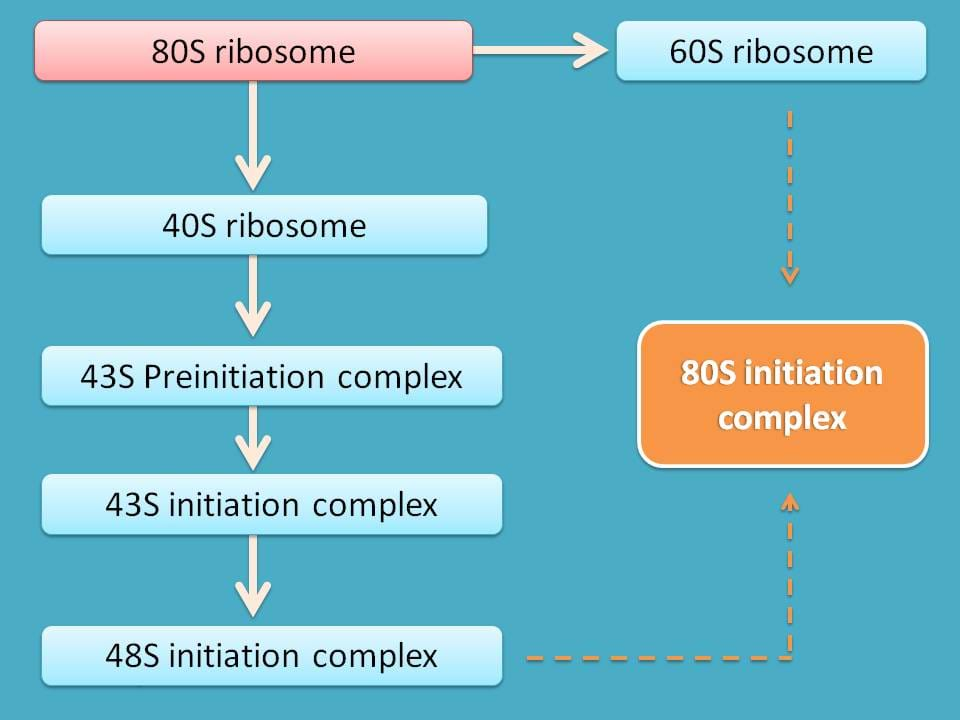
60S and 40S subunits
So the first step in translation of proteins is the separation of 80S ribosome into 60S and 40S subunits.
As you know, when 80S ribosome is dissociated into 60S and 40S subunits, they have equal tendency to re-join and bring back to single 80S ribosome.
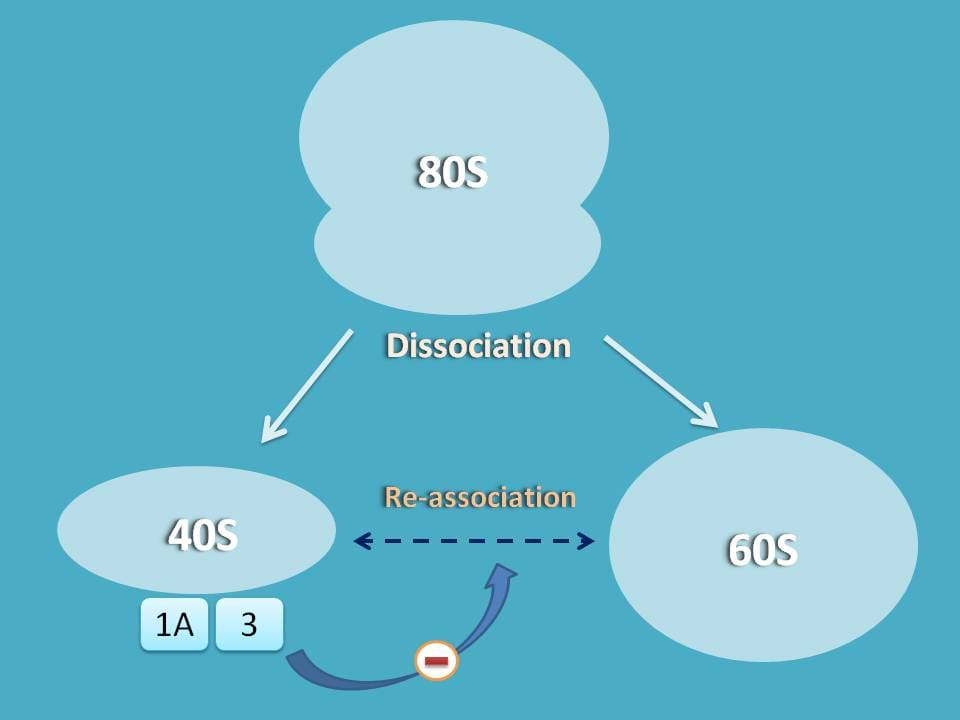
So, we should have some mechanism that stabilizes the dissociation and prevents re-association of these subunits.
Here initiation factors come into the play. Since we are discussing protein translation in eukaryocytes, they are termed as eukaryocytic initiation factors (eIF).
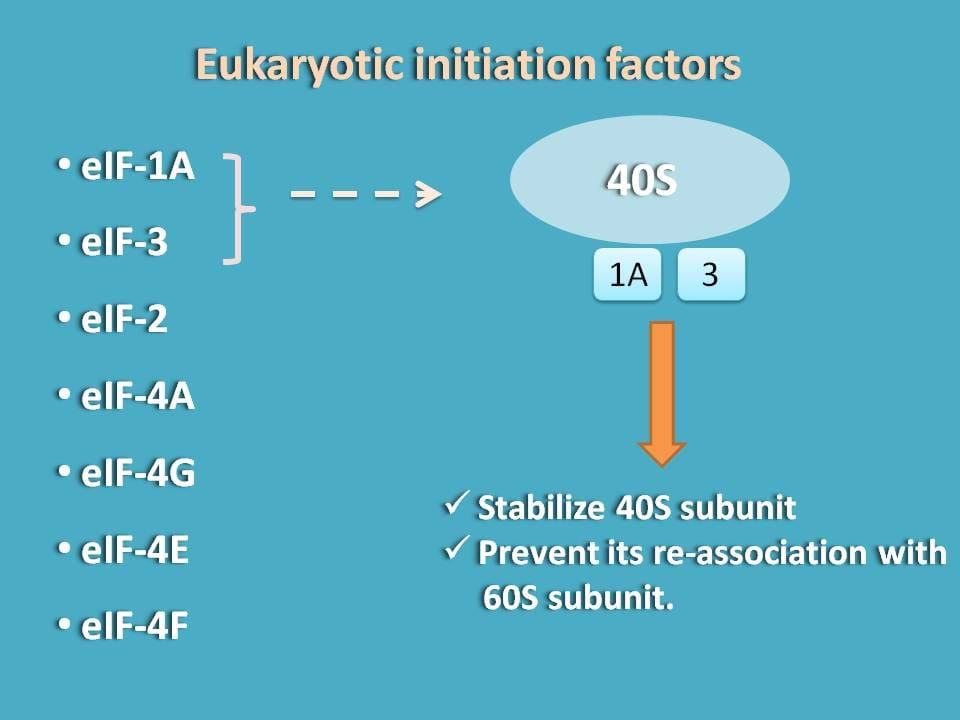
The first two initiation factors that stabilise 40S subunit are eIF-3 and eIF-1A. Once they bind to 40S subunit they stabilise and reduce its interaction with 60S subunit.
43S preinitiation complex
Okay, ribosome is dissociated and now 40S subunit is ready for modification. Initially, it should be added with tRNA which brings the initiator amino acid for translation.
What is that initiator amino acid?
It is simply methionine, which is supplied as aminoacyl tRNA that is required for initiation of translation. Synthesis of methionine aminoacyl tRNA is mediated by aminoacyl synthetase enzyme utilising one ATP molecule.
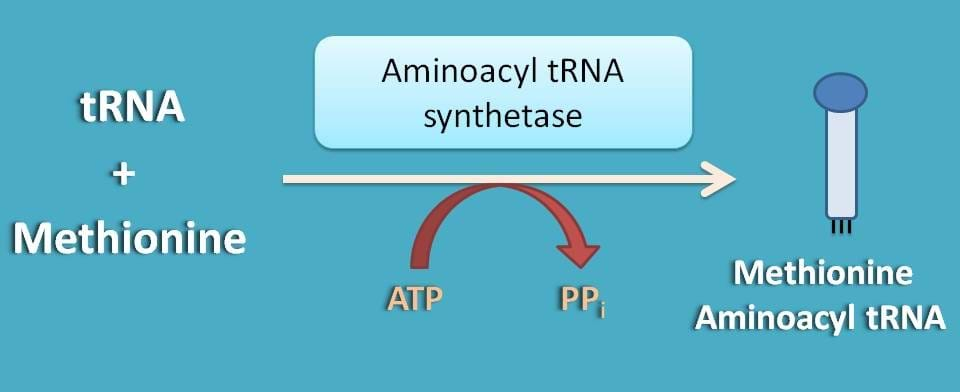
This methionine aminoacyl tRNA (met-tRNA) can form a ternary complex with initiation factor 2 and GTP which is then attached to 40S subunit to form first preinitiation complex 43S.
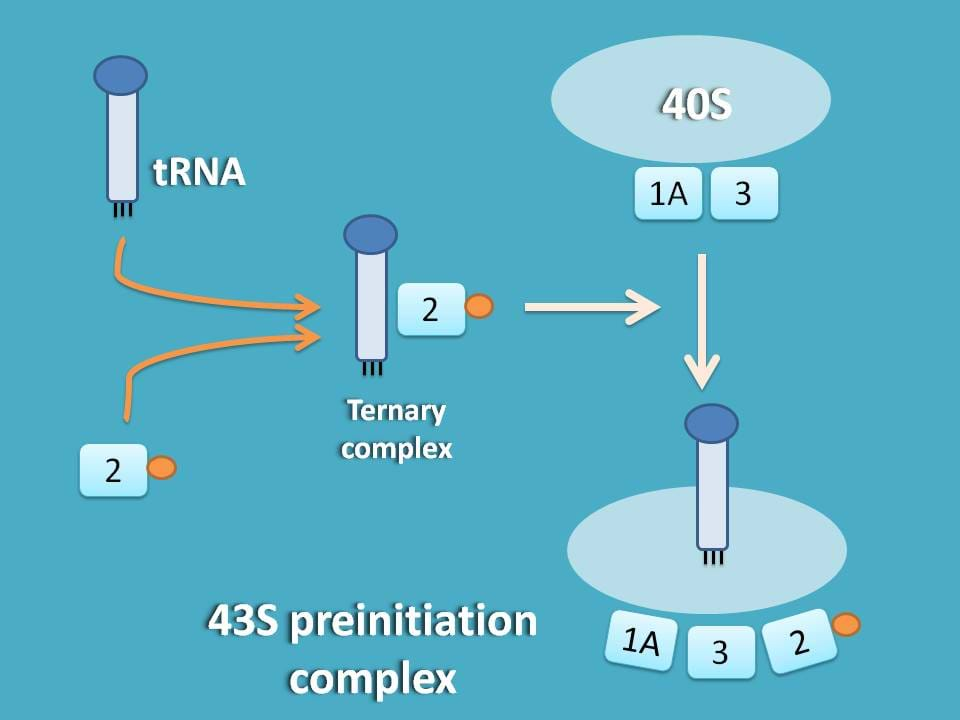
Remember here that the same factors such as eIF-3 and eIF-2A which are responsible for stabilisation of 40S subunit are also responsible for formation f 43S preinitiation complex.
43S initiation complex
Now, the next step is to bind mRNA to 43S preinitiation complex.
Here we have two problems.
First, mRNA is generally capped at 5’ end which should be activated in order to bind to 43s ribosome complex.
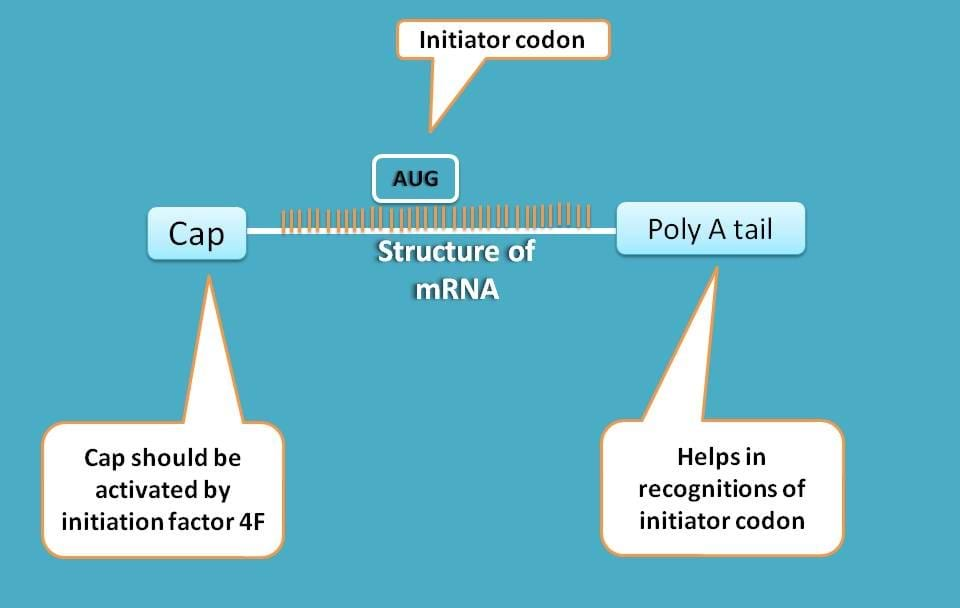
Second, the secondary structure of mRNA is highly complex which should be simplified for translation to proceed.
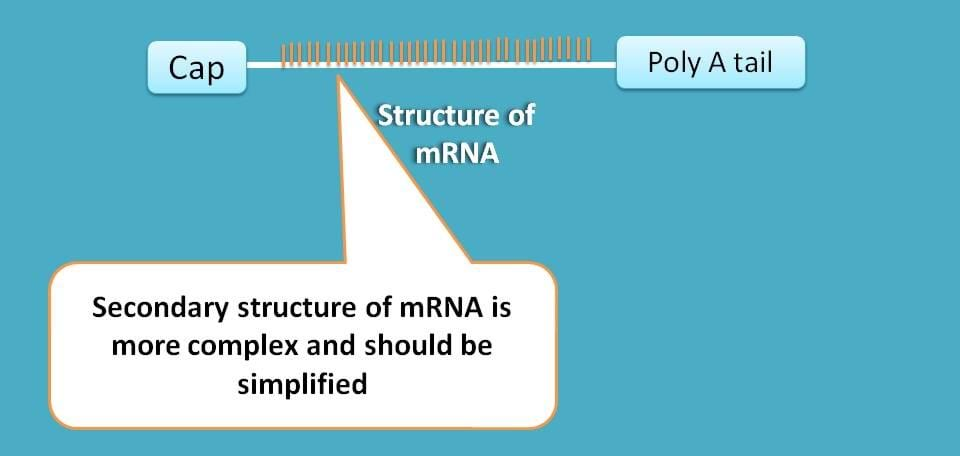
Fortunately we have again specific initiation factors for solving these two problems.
Few of the factors like eIF-4E, eIF-4G and eIF-4A combine to form eIF-4F. Now this eIF-4F binds to cap on mRNA and helps in binding it to 43S preinitiation complex.
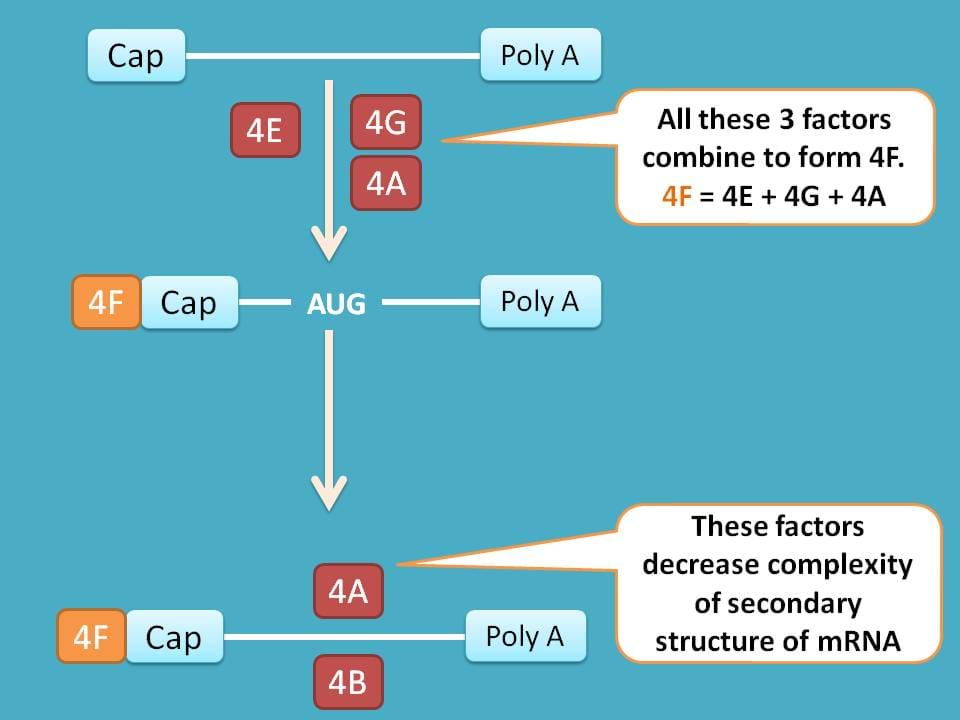
Similarly factors like eIF-4A and eIF-4B reduce the complexity of mRNA by modifying their secondary structure.
img
Now, the resulting mRNA binds to 43S preinitiation complex and converted into 43S initiation complex.
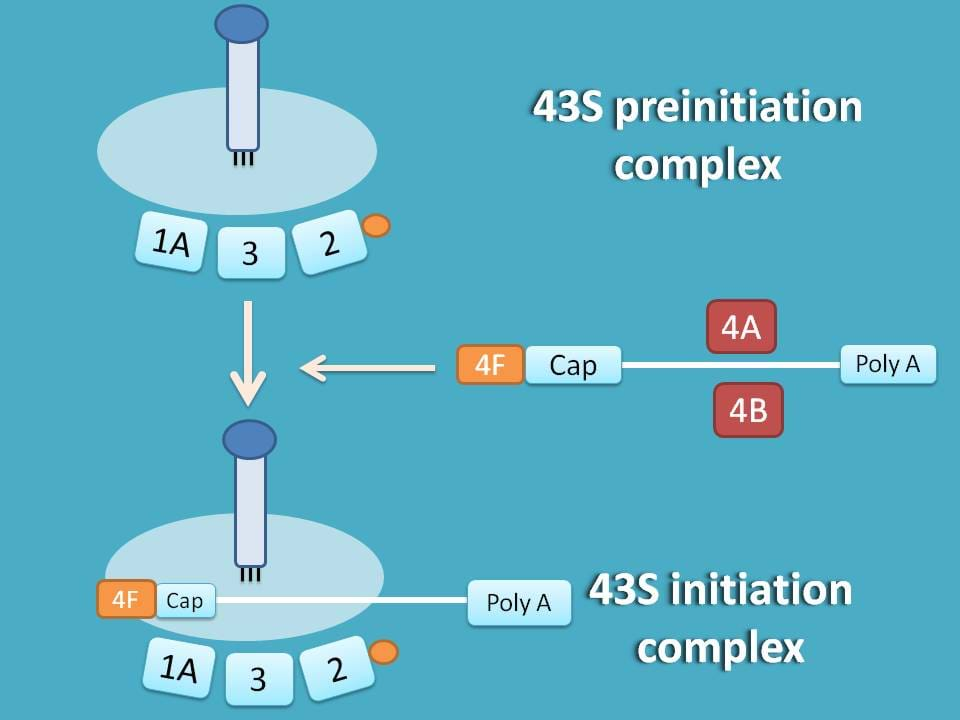
48S initiation complex
Okay, we have mRNA attached to 43S ribosome, but where to start the translation?
So, next step is to identify initiation point on mRNA. As you know methionine acts as initiator for protein synthesis coded by AUG, this codon sequence should be identified on the mRNA.
43S initiation complex formed above immediately scans from 5’end for specific sequence containing AUG.
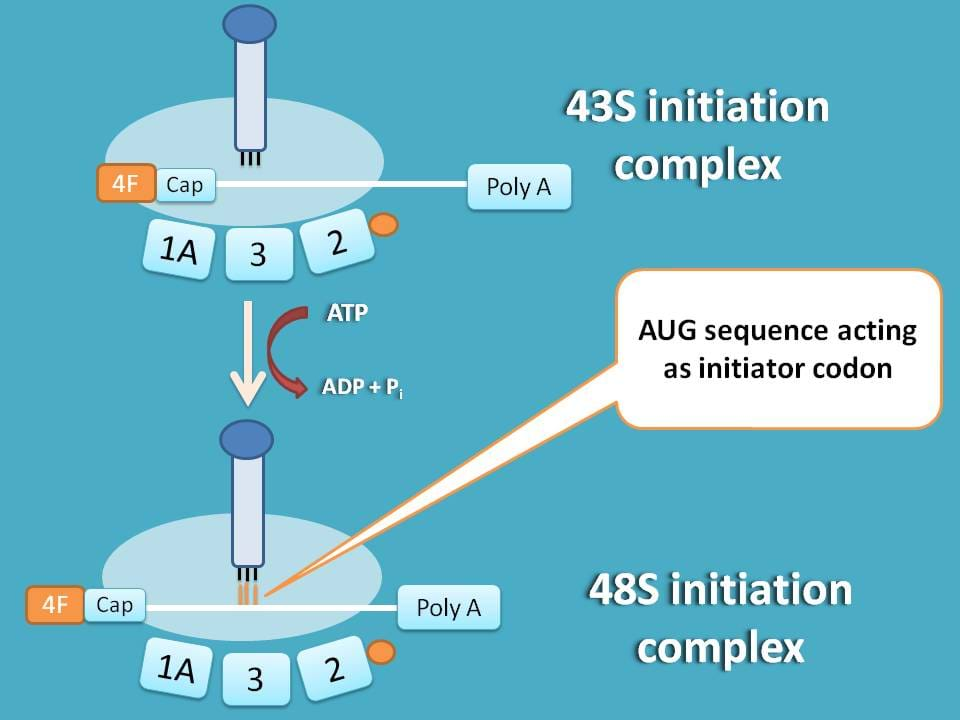
You can see here that mRNA is shifted to left to match with methionine tRNA with codon AUG.
80S initiation complex
This is the final step in initiation step. 48S initiation complex combines with 60S subunit to produce 80S initiation complex.
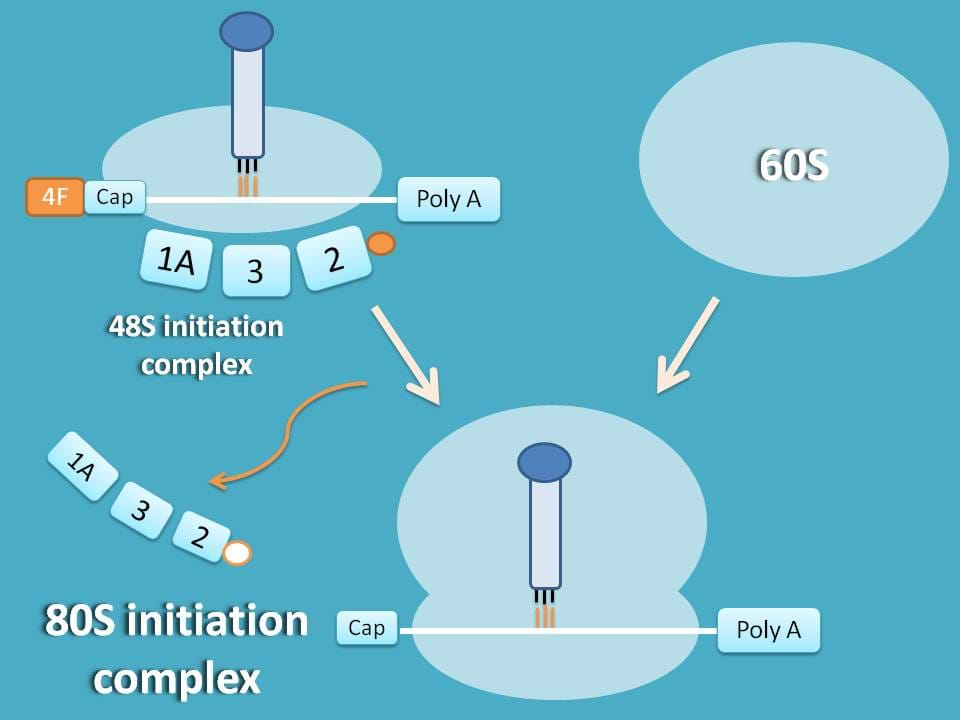
At this step, factors like eIF-1A, eIF-2 and eIF 3 are released and reused for next cycle of initiation.
A little difference exists at eIF-2 which is released as GDP associated and therefore reconverted into GTP associated factor by another factor eIF-2B.
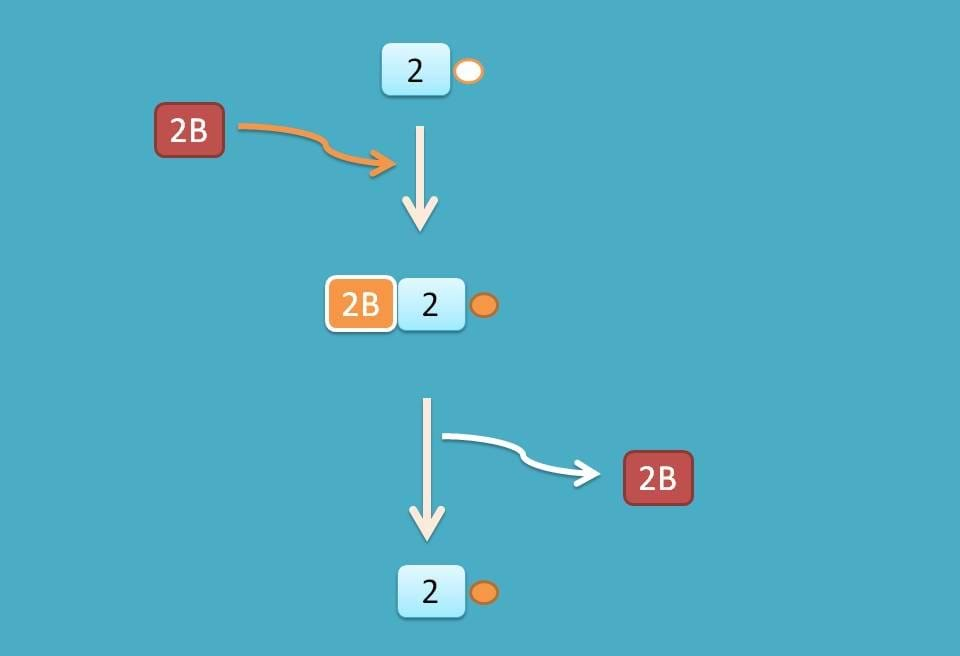
Now ribosome with active mRNA is available for protein translation. During this development, three binding sites are created on 60S subunit commonly known as A, P and E sites.
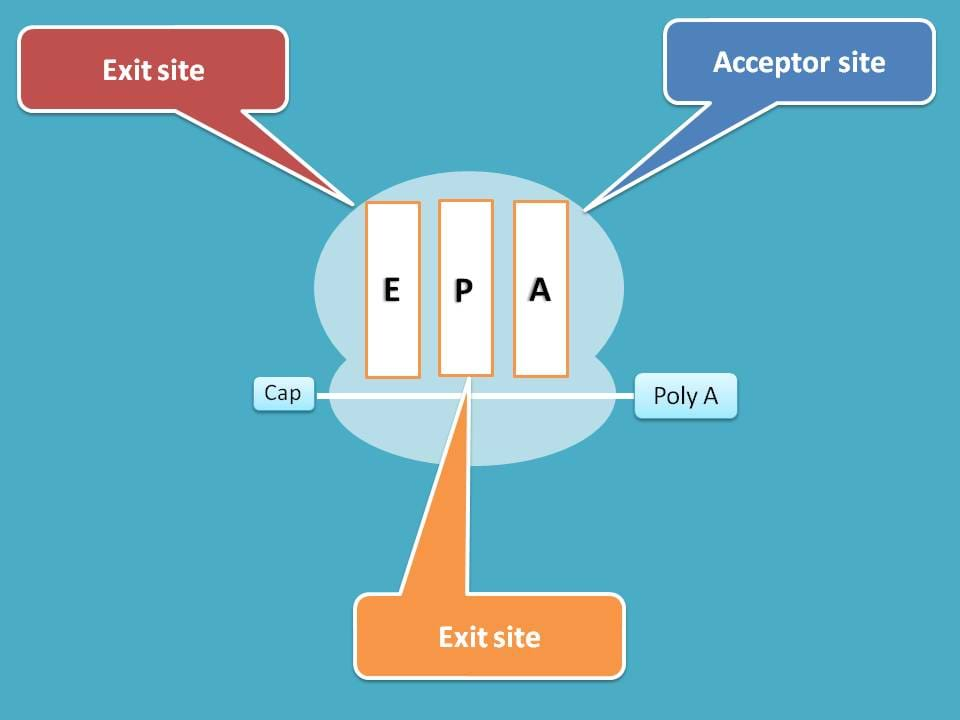
A site is the acceptor site for incoming aminoacyl tRNA, P site the peptide site where peptide chain is elongated and E site is the exit site for removal vacant tRNA.
You can observe here that, P site already has one tRNA with amino acid methionine which acts as initiator for protein translation.
That’s it. Now initiation step is over and everything is set ready for elongation step.
So, let’s continue our discussion with elongation step.
Elongation
Here real translation process takes place to prepare polypeptide chain. Again here three steps take place continuously in a cyclic fashion to synthesize peptide chain. They are
- Attachment of tRNA
- Transpeptidation
- Translocation
Let’s see these steps in detail.
Attachment of tRNA
Just like in the initiation step, here again each amino acid is carried by tRNA in the form of aminoacyl tRNA.
Where it attaches?
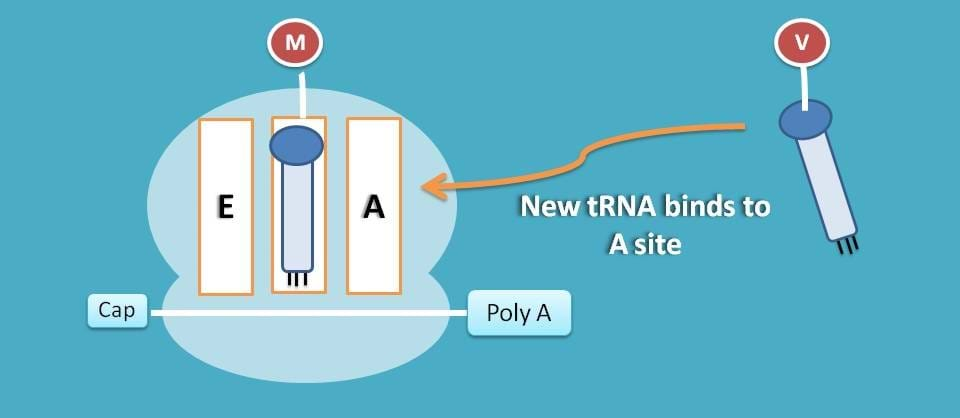
As you have seen earlier, P site is already occupied therefore new tRNA is attached to A site.
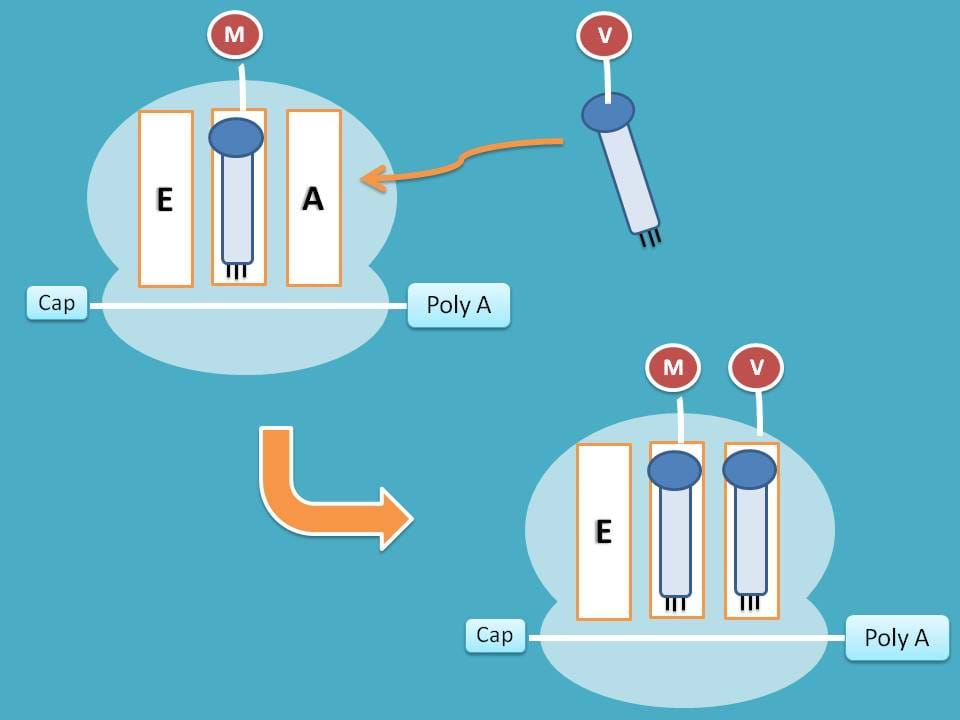
This attachment is activated by eukaryotic elongation factor 1α (eEF-1α) and this step consumes one GTP molecule.
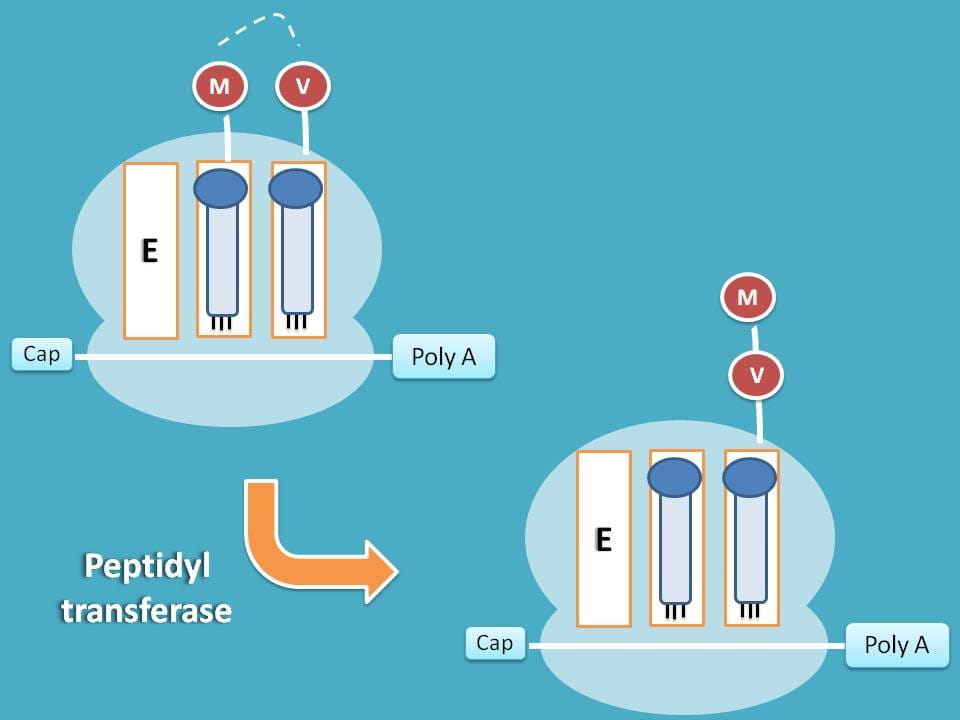
Transpeptidation
Now both A and P sites have tRNA each attached with amino acid. In order to prepare peptide chain these amino acids should be combined. This can be achieved by transfer of amino acid from P site to that on to A site and simultaneous peptide bond formation.
This step is carried by peptidyl transferase enzyme.
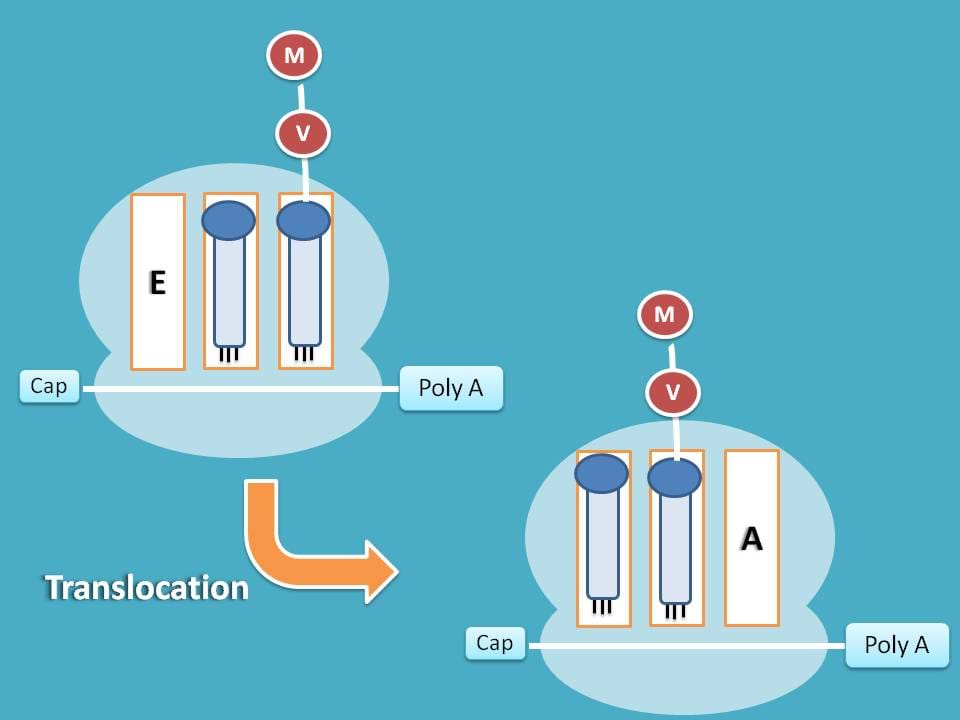
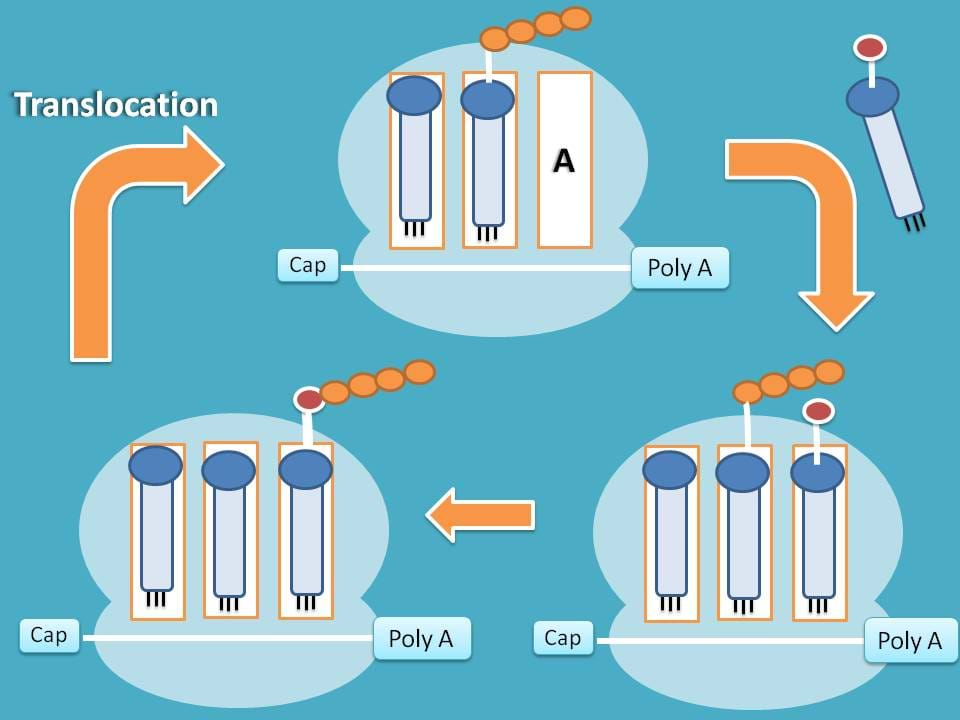
Translocation
Finally, the tRNA carrying peptide chain at A site is shifted to P site giving way to new tRNA for attachment at A site.
Note: Each vacant tRNA is shifted to E site and the removed from the ribosome as translation proceeds.
In this way, the three steps in elongation are repeated several times with each cycle adding one amino to growing polypeptide chain.
Termination
There should be an end to any process so to the translation. But how mRNA can give signal for that?
As you are familiar, few of the codons doesn’t code for any amino acid but just send signal for termination of translation. These codons such as UAA, UGA and UAG are called as stop codons which stop translation.

When codons in mRNA reach to these codons, simply a water molecule is added to peptide chain and chain is released.
This release of peptide chain is facilitated by eukaryotic releasing factor (eRF).
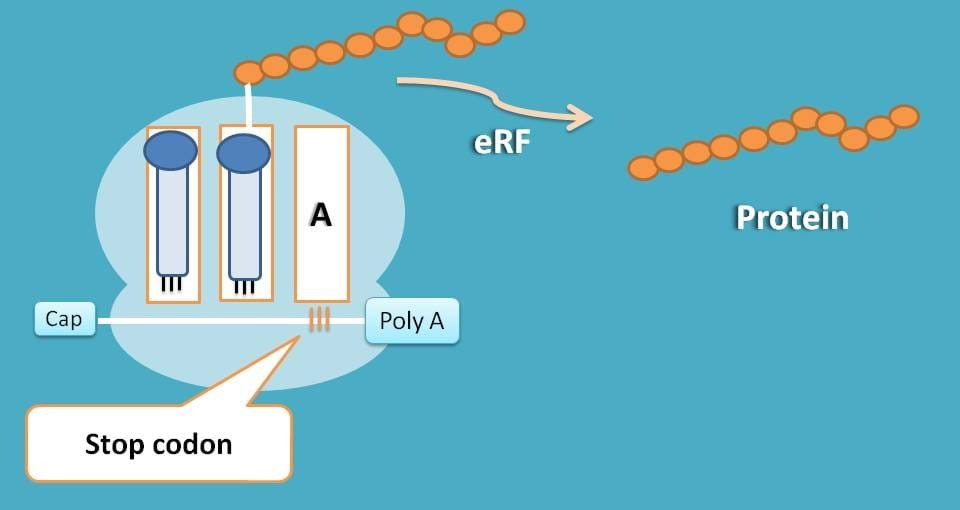
That’s the story. Now each peptide synthesized at a particular sequence and cleaved at specific length act as a unique protein either building the cellular structures or managing the cellular functions.
Hope you have enjoyed reading this article. Post your comments and don’t forget to share this with your friends.